Activity 8: Classification predictions and gradients Live#
2026-02-24
Imports and data loading#
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from palmerpenguins import load_penguins
from sklearn.linear_model import LogisticRegression
# Load Palmer Penguins and filter to Adelie vs Chinstrap
df = load_penguins()
penguin_df = df[df["species"].isin(["Adelie", "Chinstrap"])].dropna(subset=["bill_length_mm", "bill_depth_mm"])
# Features: bill_length_mm and bill_depth_mm
X_train = penguin_df[["bill_length_mm", "bill_depth_mm"]].values
feature_names = ["bill_length_mm", "bill_depth_mm"]
# Labels: Adelie = 0, Chinstrap = 1
y_train = (penguin_df["species"] == "Chinstrap").astype(int).values
print(f"Number of examples: {len(y_train)}")
print(f"Number of Adelie (y=0): {np.sum(y_train == 0)}")
print(f"Number of Chinstrap (y=1): {np.sum(y_train == 1)}")
print(f"Feature matrix shape: {X_train.shape}")
penguin_df['y'] = y_train
Number of examples: 219
Number of Adelie (y=0): 151
Number of Chinstrap (y=1): 68
Feature matrix shape: (219, 2)
Part 1: Linear decision boundaries#
The above cell loads in our data into penguin_df.
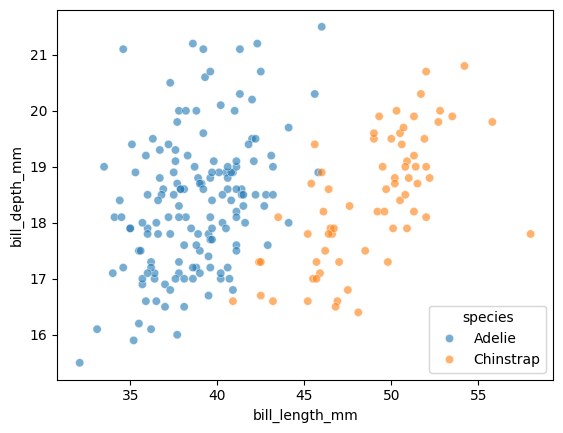
Let’s visualize our two features, colored by species. Let’s create a scatterplot with:
x="bill_length_mm"y="bill_depth_mm"hue="species"data=penguin_dfalpha=0.6
sns.scatterplot(
x="bill_length_mm",
y="bill_depth_mm",
hue="species",
data=penguin_df,
alpha=0.6
)
display(penguin_df.head())
| species | island | bill_length_mm | bill_depth_mm | flipper_length_mm | body_mass_g | sex | year | y | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | Adelie | Torgersen | 39.1 | 18.7 | 181.0 | 3750.0 | male | 2007 | 0 |
| 1 | Adelie | Torgersen | 39.5 | 17.4 | 186.0 | 3800.0 | female | 2007 | 0 |
| 2 | Adelie | Torgersen | 40.3 | 18.0 | 195.0 | 3250.0 | female | 2007 | 0 |
| 4 | Adelie | Torgersen | 36.7 | 19.3 | 193.0 | 3450.0 | female | 2007 | 0 |
| 5 | Adelie | Torgersen | 39.3 | 20.6 | 190.0 | 3650.0 | male | 2007 | 0 |
Suppose that we have the following weights:
# Weights for a 2D model using bill_length_mm and bill_depth_mm
w0 = -24.5
w1 = 1.35 # weight for bill_length_mm (x1)
w2 = -1.9 # weight for bill_depth_mm (x2)
We can again solve for the decision boundary by setting \(h=0\):
\(x_2\) is on the y-axis in our plot, so we can solve for it:
Let’s plot this line over our data:
# Two x1 endpoints to draw the boundary line
x1 = np.array([35, 55])
# TODO compute the corresponding x2 values on the decision boundary
x2 = -(w0 + w1*x1) / w2
sns.scatterplot(
x="bill_length_mm",
y="bill_depth_mm",
hue="species",
data=penguin_df,
alpha=0.6
)
plt.plot(x1, x2, color="black")
plt.ylim(15,22)
#plt.xlim(40, 50)
(15.0, 22.0)
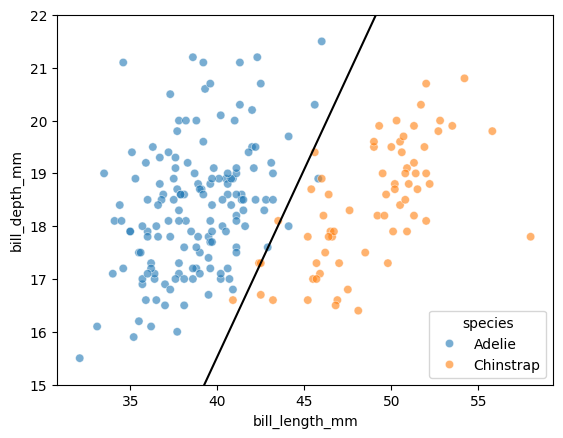
Takeaway
Even though the sigmoid function is non-linear, the decision boundary is going to be linear in the features!
Part 2: Generating probabilities and predictions#
LogisticRegression from scikit-learn works the same way as LinearRegression: we call .fit(X, y) to train the model.
# TODO creates a LogisticRegression model and fit it on X and y
model = LogisticRegression()
model.fit(X_train, y_train)
# Print the learned weights
print(f"Intercept (w0): {model.intercept_[0]:.4f}")
print(f"Weights: {model.coef_[0]}")
print(f" w1 (bill_length): {model.coef_[0][0]:.4f}")
print(f" w2 (bill_depth): {model.coef_[0][1]:.4f}")
Intercept (w0): -24.5494
Weights: [ 1.35804435 -1.92413248]
w1 (bill_length): 1.3580
w2 (bill_depth): -1.9241
But unlike LinearRegression, logistic regression can output probabilities using the .predict_proba() method:
This returns an array of shape (n, 2) where:
Column 0: probability of class 0 (Adelie)
Column 1: probability of class 1 (Chinstrap)
# TODO: call predict_proba() on the model with input X
proba = model.predict_proba(X_train)
print(f"Shape of proba: {proba.shape}")
print(f"\nFirst 5 rows of proba:")
print(proba[:5])
Shape of proba: (219, 2)
First 5 rows of proba:
[[9.99407822e-01 5.92177979e-04]
[9.87709002e-01 1.22909979e-02]
[9.88508221e-01 1.14917793e-02]
[9.99992825e-01 7.17502671e-06]
[9.99979912e-01 2.00882817e-05]]
Predictions are then made by picking the class with the highest probability:
# TODO: call predict on the model with input X
preds = model.predict(X_train)
print(f"Shape of preds: {preds.shape}")
print(f"\nFirst 5 rows of preds:")
print(preds[:5])
Shape of preds: (219,)
First 5 rows of preds:
[0 0 0 0 0]
Let’s look at how to compute these outputs on our own.
Suppose we have the \(P(y = 1)\) probabilities from the slides:
proba1_chinstrap = np.array([0.16, 0.78, 0.29, 0.92, 0.52])
print(proba1_chinstrap)
print(proba1_chinstrap.shape)
[0.16 0.78 0.29 0.92 0.52]
(5,)
To convert a 1D array of shape (n,) into a 2D array with shape (n, 1), we can use np.reshape:
# the -1 means "infer the size of the other dimension"
proba1_chinstrap = proba1_chinstrap.reshape(-1, 1)
print(proba1_chinstrap)
print(proba1_chinstrap.shape)
[[0.16]
[0.78]
[0.29]
[0.92]
[0.52]]
(5, 1)
Since we know that probabilities must sum to 1, what is the line of code that can generate the 2D array for the \(P(y = 0)\) probabilities? Hint: recall what happens when we perform arithmetic between a number and a NumPy array.
Submit the line of code here: https://pollev.com/tliu
# TODO: this can be done in one line!
proba0_adelie = 1 - proba1_chinstrap
print(proba0_adelie)
[[0.84]
[0.22]
[0.71]
[0.08]
[0.48]]
np.column_stack() then can be used to stack arrays column-wise:
proba_array = np.column_stack([proba0_adelie, proba1_chinstrap])
print(proba_array)
[[0.84 0.16]
[0.22 0.78]
[0.71 0.29]
[0.08 0.92]
[0.48 0.52]]
How do we then go from this proba_array to a 1D array of predictions?
np.argmax() finds the index of the maximum value, and can be used with axis operations
axis=1 means: find the index of the maximum value within each row
axis=0 means: find the index of the maximum value within each column
no axis means: find the index of the maximum value in the entire array
Note the shape changes:
print(proba_array)
print(proba_array.shape) # (n, 2)
print("-----------------")
print(np.argmax(proba_array, axis=1))
[[0.84 0.16]
[0.22 0.78]
[0.71 0.29]
[0.08 0.92]
[0.48 0.52]]
(5, 2)
-----------------
[0 1 0 1 1]
What axis should we use to generate predictions from proba_array?
predictions = np.argmax(proba_array, axis=1)
Part 3: Broadcasting practice#
On HW 1, we computed the gradient descent update rule “manually” for each of the weights:
Today, we’ll practice using broadcasting to perform a gradient update for all the weights at once.
Suppose we had some loss function \(\mathcal{L}\) such that the gradient with respect to weights \(w_1, w_2, \ldots, w_p\) is given by:
This gives the gradient update rule for all the weights:
Given the y’s as a 1D array y of shape (n,), and the data matrix X of shape (n, p), we can compute the gradient update for all the weights at once using broadcasting.
Let’s trace through the broadcasting with a small example (\(n=3\) examples, \(p=2\) features):
# Small example to show broadcasting
X = np.array([[1, 2],
[2, 4],
[5, 6]])
y = np.array([1, 0, 1])
print(f"X shape: {X.shape}")
print(X)
print("-----------------")
print(f"y shape: {y.shape}")
print(y)
X shape: (3, 2)
[[1 2]
[2 4]
[5 6]]
-----------------
y shape: (3,)
[1 0 1]
Out goal is to compute the summation gradient term for all the weights simultaneously with broadcasting:
To do so, we multiply each feature of X by y element-wise:
And then sum over the rows:
In order for y to broadcast with X, we need to reshape y to have shape (3, 1):
# TODO: update y to be 2D with shape (n, 1)
y_reshaped = y.reshape(-1, 1)
print(y_reshaped)
print(y_reshaped.shape)
[[1]
[0]
[1]]
(3, 1)
# this will give us a broadcasting error
#y * X
# TODO: use y_shaped to properly broadcast with X
y_reshaped * X
array([[1, 2],
[0, 0],
[5, 6]])
Then, we use axis=0 to np.sum over the rows:
# TODO take the sum over the rows, resulting in shape (p,)
# if this array is (n,p)
result = np.sum(y_reshaped * X, axis=0)
print(result)
print(result.shape)
[6 8]
(2,)
result is a 1D array with shape (p,), which can be used to update all the weights at once:
w_old = np.array([0, 1])
alpha = 0.1
# TODO: all 2 weights are updated at once without a loop
w_new = w_old - alpha * result
NumPy broadcasting allows this to work for any number of features \(p\), and the gradient update on HW 2 will use this same technique.
